Elucidation of the molecular mechanisms by which plants respond to osmotic stress elicited by desiccation, salt, and cold is required to increase stress tolerance in crops. Many physiological responses to ensure acclimation adverse environmental conditions require the synthesis and perception of the plant hormone abscisic acid (ABA).
We have demonstrated that if a germinated seedling is challenged with water deficit, it can arrest its growth until adequate water status is restored. This growth arrest to ensure seedling survival is dependent on ABA and requires the transcription factor ABI5. In the absence of ABA, ABI5 is targeted for ubiquitin-mediated proteolysis. A screen for interacting partners of ABI5 led to the identification of AFP, which enhances ABI5 degradation in the absence of ABA.
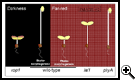 Reproduced, with permission, from Genes and Development Vol 15 (14). © 2001 by Cold Spring Harbor Laboratory Press.
Reproduced, with permission, from Genes and Development Vol 15 (14). © 2001 by Cold Spring Harbor Laboratory Press.The ubiquitin-like protein SUMO also participates in ABA signal transduction. We demonstrated that increased SUMOylation attenuates ABA-mediated growth inhibition and amplified induction of certain ABA-and stress-responsive genes. Accordingly, a reduction in the expression of an Arabidopsis SUMO-conjugase accentuates ABA-mediated growth inhibition.
Another aspect of our research into ABA-mediated molecular responses to stress concerns assessment of the roles of microRNAs in mounting adaptive responses to water deprivation. In collaboration with members of Terry Gaasterland's lab at Rockefeller University, we have used an in silico approach to screen for potential microRNA genes in the Arabidopsis genome and are investigating several candidates likely to mediate post-transcriptional regulation of ABA-mediated responses to water deprivation.
References:
Zhang X, Garreton V, Chua N-H (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev. 19:1532-1543
Lopez-Molina L, Mongrand S, Kinoshita N, Chua NH (2003) AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev 17:410-418
Lois LM, Lima CD, Chua NH (2003) Small ubiquitin-like modifier modulates abscisic acid signaling in Arabidopsis. Plant Cell 15: 1347-1359
Duque P, Chua NH (2003) IMB1, a bromodomain protein induced during seed imbibition, regulates ABA- and phyA-mediated responses of germination in Arabidopsis. Plant J 35:787-799
Wu Y, Sanchez JP, Lopez-Molina L, Himmelbach A, Grill E, Chua NH (2003) The abi1-1 mutation blocks ABA signaling downstream of cADPR action. Plant J 34:307-315
Lopez-Molina L, Mongrand B, McLachlin DT, Chait BT, Chua NH (2002) ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J 32:317-328
Hoth S, Morgante M, Sanchez JP, Hanafey MK, Tingey SV, Chua NH (2002) Genome-wide gene expression profiling in Arabidopsis thaliana reveals new targets of abscisic acid and largely impaired gene regulation in the abi1-1 mutant. J Cell Sci 115:4891-4900
Lopez-Molina L, Mongrand S, Chua NH (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci USA 98:4782-4787
Lemichez E, Wu Y, Sanchez JP, Mettouchi A, Mathur J, Chua NH (2001) Inactivation of AtRac1 by abscisic acid is essential for stomatal closure. Genes Dev 15:1808-1816
Sanchez JP, Chua NH (2001) Arabidopsis PLC1 is required for secondary responses to abscisic acid signals. Plant Cell 13:1143-1154